Intelligent Bioinformatics for Every Team: Self-Optimizing Compute, Co-Scientist, and Autonomous Research
Today, at the Boston Nextflow Summit, I presented on how the leading scientific teams are orchestrating discovery in the AI era. We're launching GPU metrics, Intelligent Compute (smarter cloud scheduling), Co-Scientist (interactive research intelligence), and a demonstration of how this all comes together for autonomous experiments. This post walks through everything we launched and what it means for the teams doing science with Nextflow today — and for the agents who will run them tomorrow.
The AI Acceleration is Compounding
Scientific discovery is accelerating, and the future of computing behind it isn't just agentic. New biological models that predict sequence to function, structural predictions and entire virtual cells are reshaping how insights can be generated. We’re already seeing impact: the first AI-discovered drug cleared Phase IIa, AI-guided patient selection showed a 15% improvement in survival risk, and 15–20 AI-originated drugs are expected to enter Phase III in 2026.
At the agentic layer, frontier models are able to take on ever larger tasks from designing experiments to autonomous execution. Every process in the life sciences is being redesigned with AI.
Yet the fundamentals of science haven't changed. Keeping the work we do FAIR is the top priority whether the research involves agentic systems or not. We've spent over a decade solving these problems for bioinformatics. Now we're applying our expertise as we build Seqera into the intelligent engine for AI-powered science. And the foundations that enable verification and reproducibility today become even more important.
The Rise of the GPU
As GPUs become central to biological models, teams need the same visibility into GPUs that they've always had for CPU workloads. GPU metrics are now first-class citizen in Seqera. Every task that uses a GPU now records type, driver version, utilization, and average and peak memory, all visible in task detail logs and metric visualization graphs. You can see exactly what GPU resources they're consuming on a per-task level across their workflows and make rightsizing decisions on that basis.
Seqera Intelligent Compute
Cloud batch schedulers were designed to replicate the HPC model, but that static approach doesn't leverage the cloud's elasticity, leading to overprovisioned resources, manual tuning, and wasted spend.
Seqera Intelligent Compute is a new-generation compute service that abstracts away the complexity of scaling pipelines in the cloud.. It continuously optimizes your cloud spend and resource utilization for more-efficient workflows, without requiring any manual configuration from your team. Intelligent Compute learns from every run and recovers from failures like preemptions and out-of-memory errors without intervention.
- →Optimized spot: Start with spot instances, automatically fall back to on-demand. Always get the best price without risking failed jobs.
- →Right-sizing: Predict what each task actually needs based on previous runs, not what was requested, so nothing is overprovisioned.
- →Job-packing: Pack tasks densely onto fewer machines based on current and future demand, reducing the total instances per run.
The result is more resource-efficient workflows: fewer instances per run, no unutilized resources, and lower overall costs. Intelligent Compute achieves levels of optimization that would take months of manual tuning to approach. Same compute, just smarter.
Seqera Co-Scientist
Seqera Co-Scientist is an intelligent, collaborative partner that brings the full power of the latest models to your scientific environment. It understands scientific context, reasons with you, and executes with intelligence, reducing the manual work and making bioinformatics accessible to everyone. Co-Scientist can work alongside you interactively or autonomously in the background with three integrated capabilities:
- →AI-native view for scientists: A simple portal that enables bench scientists to launch pipelines, manage data, and explore results independently, with guardrails that bioinformaticians and IT can configure and evolve over time.
- →Interactive research intelligence: Build, validate, optimize, and debug pipelines through CLI, IDE, MCP, or web with full workspace context, working alongside your internal AI tools and LLMs.
- →Automation with agents (coming soon): Trigger pipelines, detect and self-fix failures, and autonomously improve code with 24/7 agents. Always on, always working.
Built for Enterprise bioinformatics, Co-Scientist provides deep integration with your data, compute, and internally model with BYOK as well as complete visibility across your entire bioinformatics stack. Co-Scientist for Enterprise is model agnostic, so you can swap models without changing workflows, and it can be self-hosted within your cloud and firewall. The same intelligence is available wherever your teams work, whether that's the web, CLI, or IDE. With Co-Scientist, teams can lean into productivity without compromising on reproducibility or security, all within guardrails you control.
Autonomous Optimization
In a world where huge results can come from the smallest of changes, each decision we take, from bench to insights, steers the impact we make. Often though, the way we process our data is set in stone. Default parameters. The standard tools. The usual steps. And it makes sense because the experiment space is infinitely large. Agents change this. We are now taking advantage of the system of science that Seqera offers, to enable autonomous exploration of the vast universe of parameters, tools and code to ensure that your experiments leave no stones unturned.
Today we released results of our closed-loop optimization system built on top of Seqera. is a It autonomously optimizes combining ML optimization, agentic execution, scientific reasoning, and algorithmic rewrites to explore the experiment space and converge on better outcomes.
Improving Biomarker Discovery
We applied this to pipeline parameters to autonomously improve DIA proteomics analyses against gold-standard benchmark datasets. It proposes, runs, and ranks parameters configurations, then replans based on measured precision and recall, with no researcher intervention between runs. The result: Higher peptide recall at matched FDR.
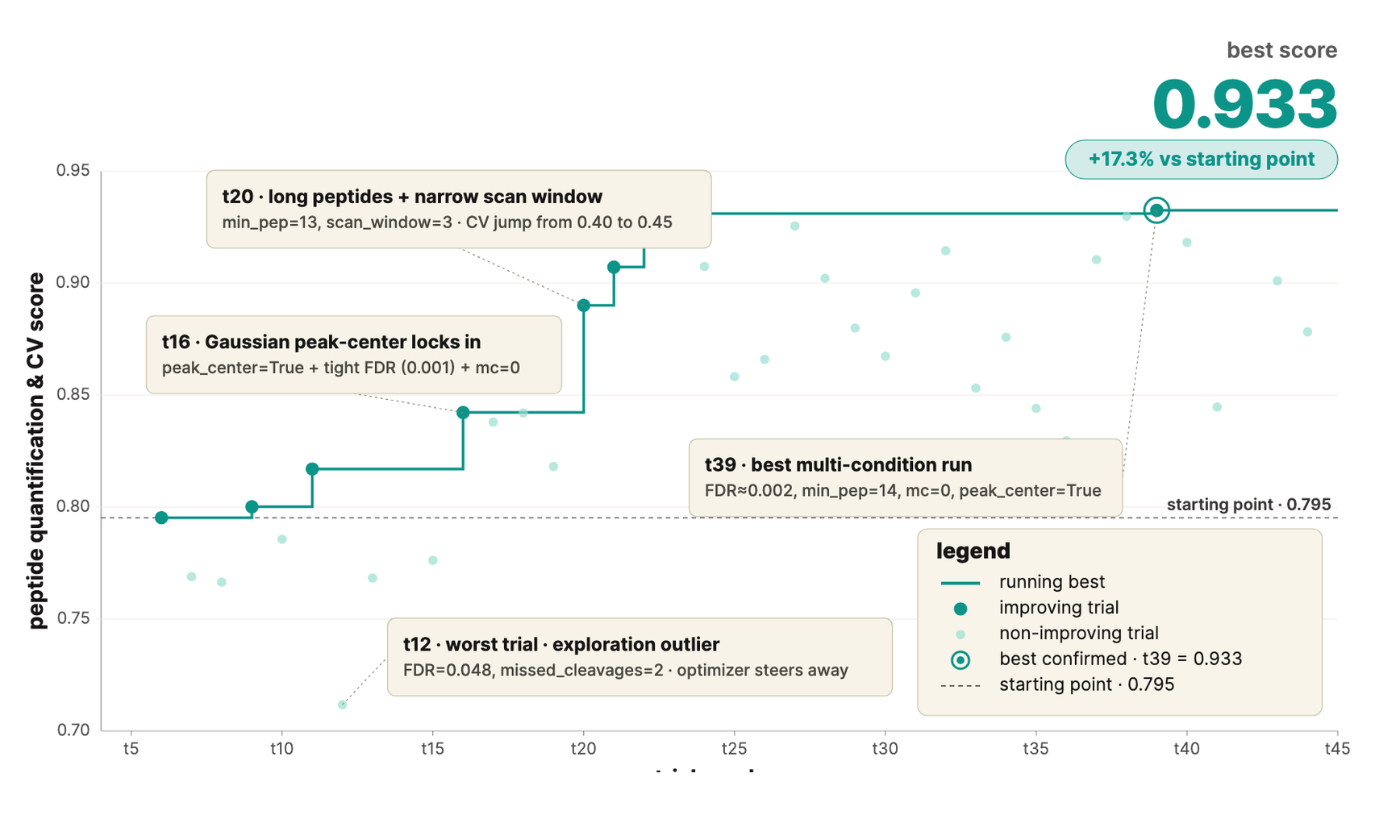
Self-Improving ML Models for Epitope Identification
The same approach can be applied to training ML models. Seqera Gradient iterates over model architecture and hyperparameters, benchmarking each round of changes against held-out ground truth. It prioritizes the configurations expected to yield the largest gains using literature search and documents the rationale in a run-lineage report. The result: precision on novel epitopes rises across iterations, without manual tuning.
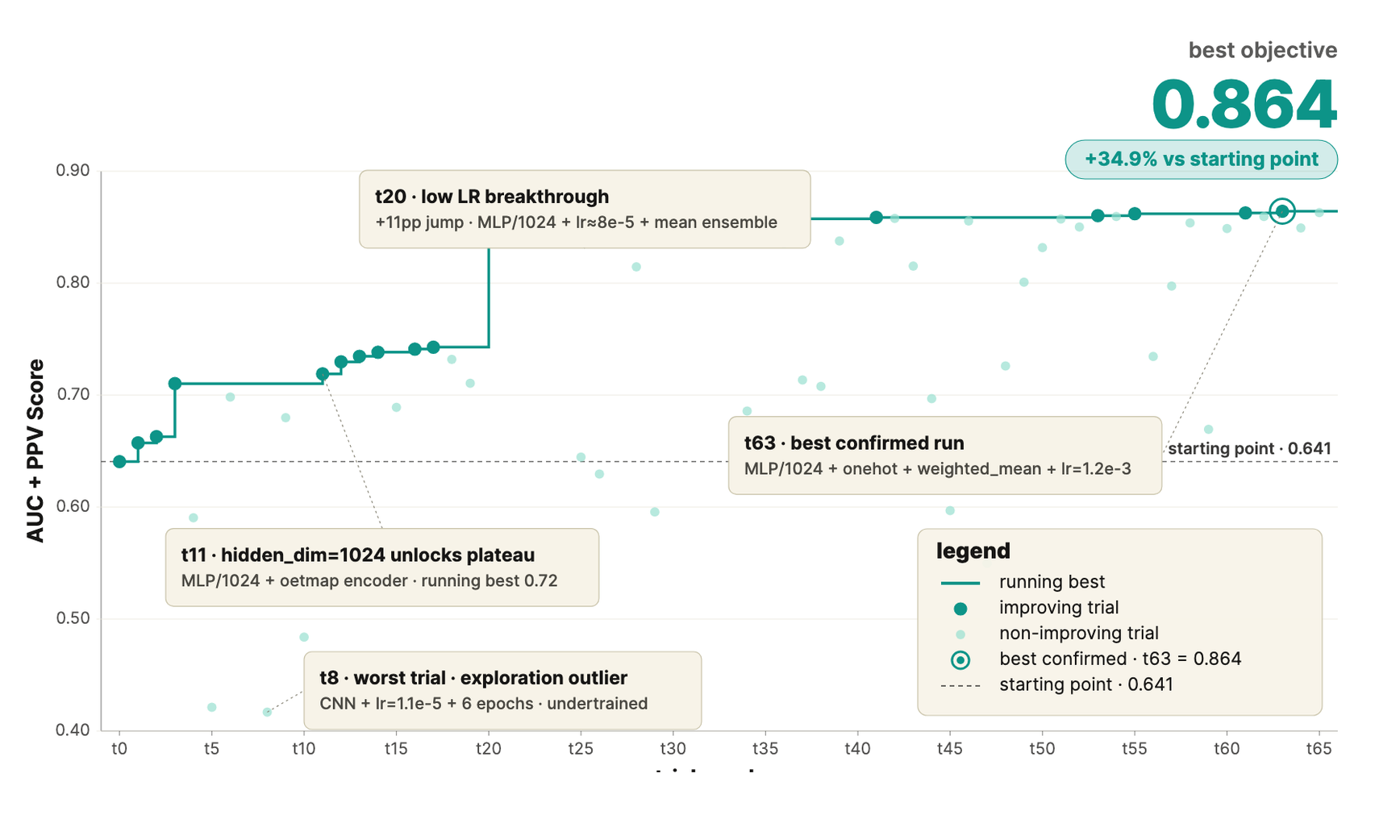
Building for What Comes Next
Everything we've shared is guided by a single conviction: the future of scientific discovery is agentic. Science is now being planned, executed, and optimized by intelligent agents working around the clock. That future demands infrastructure that is smarter, more autonomous, and more observable than anything that exists today.
Today's launch of Intelligent Compute that optimizes resources. Co-Scientist that meets every scientist where they work. And highlighting how together, Seqera closes the loop on autonomously science. These are the foundations of a system of science in AI, built on the same principles of orchestration, reproducibility, and automation that the best teams across the industry already trust. The acceleration is compounding. We cannot wait to see the impact of these advances on science
