process sayHello {
input:
val cheers
output:
stdout
script:
"""
echo $cheers
"""
}
workflow {
channel.of('Ciao', 'Hello', 'Hola') | sayHello | view()
}100% open source
High-performance data pipelines
Nextflow is the leading open-source workflow orchestrator that simplifies writing and deploying compute and data-intensive pipelines at scale on any infrastructure.
With Nextflow, scientists and bioinformaticians can easily create, share and manage reliable, reproducible scientific workflows that are portable across compute environments from desktops to clusters to clouds to serverless platforms.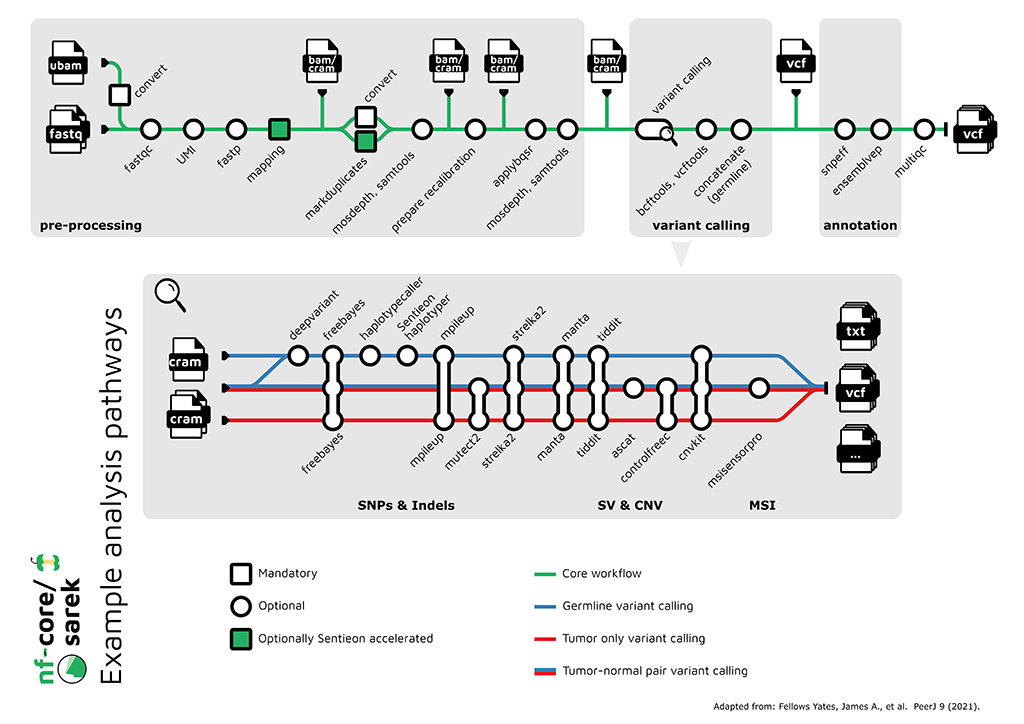
- Advanced container support for reproducible workflows
- Multiple cloud and cluster compute environments
- Source code management integrations with Git
- Massively scalable reactive dataflow programming model
- Your choice of scripting languages
Features
Simplified workflow development
Unlike other tools where pipelines can be challenging to code, deploy, and scale, Nextflow provides an expressive domain-specific language (DSL) and intuitive dataflow programming model that simplifies the development of parallel, reactive workflows. Users are free to code workflows in their language of choice using familiar tools, making it easy to adapt existing scripts to Nextflow and improving user productivity.
By using containers and abstracting compute environments from pipeline logic, Nextflow workflows are portable, scaling without modification on your choice of cloud, avoiding the risk of lock-in. Configurable retry logic and continuous checkpoints help ensure that workflows are reliable and predictable. Nextflow can automatically recover workflows and continue execution from the last completed step, avoiding unnecessary re-computation so that workflows finish on time.In good company
Trusted by the world's most innovative organizations since 2014
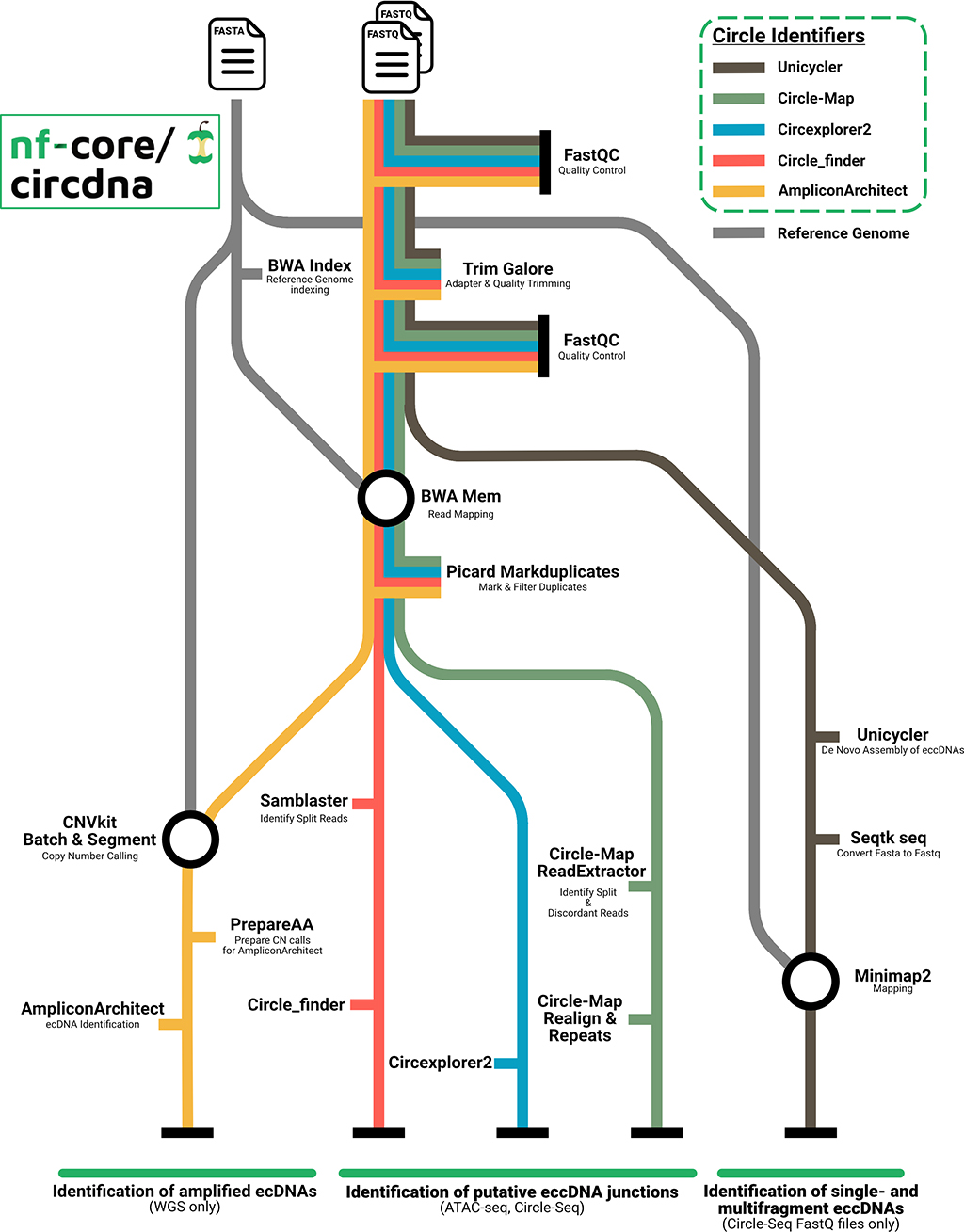
Along with a community of thousands of scientists, engineers and developers, Nextflow users can also leverage validated, high-quality open source community pipelines from organizations such as nf-core to accelerate research and significantly reduce development efforts.