The First Anniversary Release
This month we celebrate the birthday of the Tower project, introduced one year ago! 🎉🎉
A lot has changed in the world, but our mission to deliver the most seamless experience for data analysis pipelines at scale remains more vital than ever. There is now a growing community who get the vision and who are joining us on the journey.
We have been thrilled to see the project gain thousands of users and raise interest both within the Nextflow ecosystem, but also more widely. Over the past 12 months, Tower users have monitored and launched more than 110 million jobs - all with practically zero downtime!
Tower is being used extensively in the efforts to fight COVID-19 in genomic sequence analysis, drug discovery and vaccine development. Likewise, over the past months, we have begun supporting deployments of Tower within organizations and enterprises' own infrastructure, including three of the top 10 pharmaceutical companies.
Today, we are celebrating by releasing a raft of new features, including some highly anticipated community requests.
Sharing is caring
For many of us, remote work in 2020 has significantly changed our habits. While the ability to glance over a colleagues shoulder may no longer be possible, scientific collaboration should not be lost in the process. The first feature we are releasing is the ability to share your workflow executions with anyone, no matter where they are sitting.
Tower users can now say goodbye to sharing logins or tmux sessions. By selecting the share icon, a dialog allows you to specify a list of collaborators with whom the workflow is accessible. They can follow along live on the progress of the pipeline or catch up on the results at any time in the future. Collaborators are specified either by providing their username or email address, with the invited user receiving a notification email, including a link to the shared workflow.
This feature allows teams to share and replicate their analysis methods with a few clicks and provides collaborators with access to the complete pipeline execution metadata to assist in debugging and optimization.
Support for HPC batch schedulers
Today's release also adds the ability to launch your pipelines from Tower into two new computing platforms: Slurm and IBM LSF batch schedulers.
This means that you can now use Tower to deploy your Nextflow pipelines in your local Slurm or LSF HPC batch scheduler! Tower connects to the login node over SSH and submits the execution of the Nextflow pipelines as a cluster job to the queue of your choice. The Nextflow execution then continues as would any other Nextflow pipeline - with the exception it can be monitored securely from your boardroom, bedroom, or Cameroon!
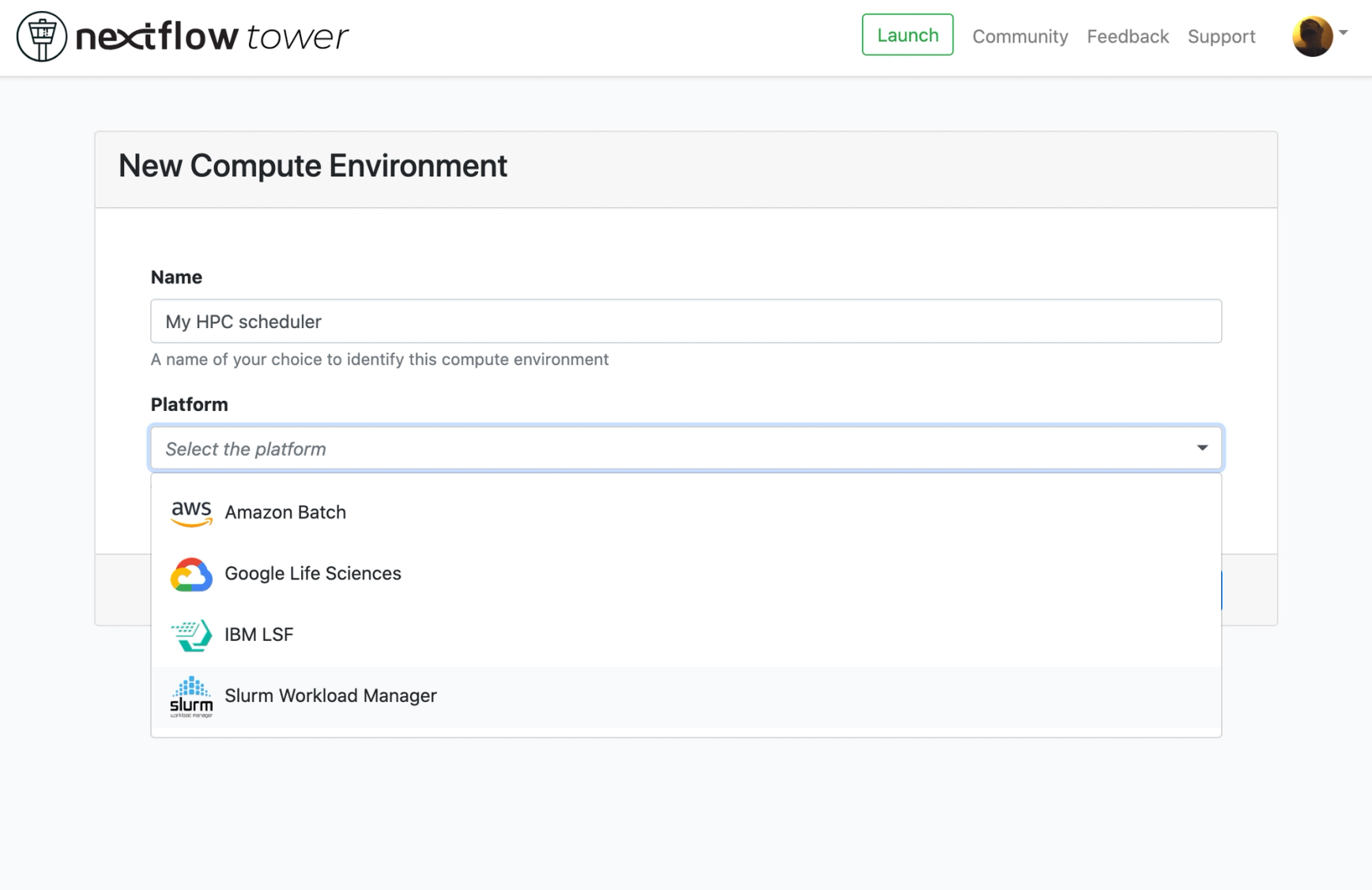
This feature greatly expands the deployment ability of Tower from cloud execution platforms such as AWS Batch and Google Life Sciences to popular batch schedulers such as Slurm and LSF connected to either on-premises and cloud resources. We are currently working towards support for other schedulers already supported by Nextflow such as Univa Grid Engine and Altair PBS Pro soon.
Google Authentication
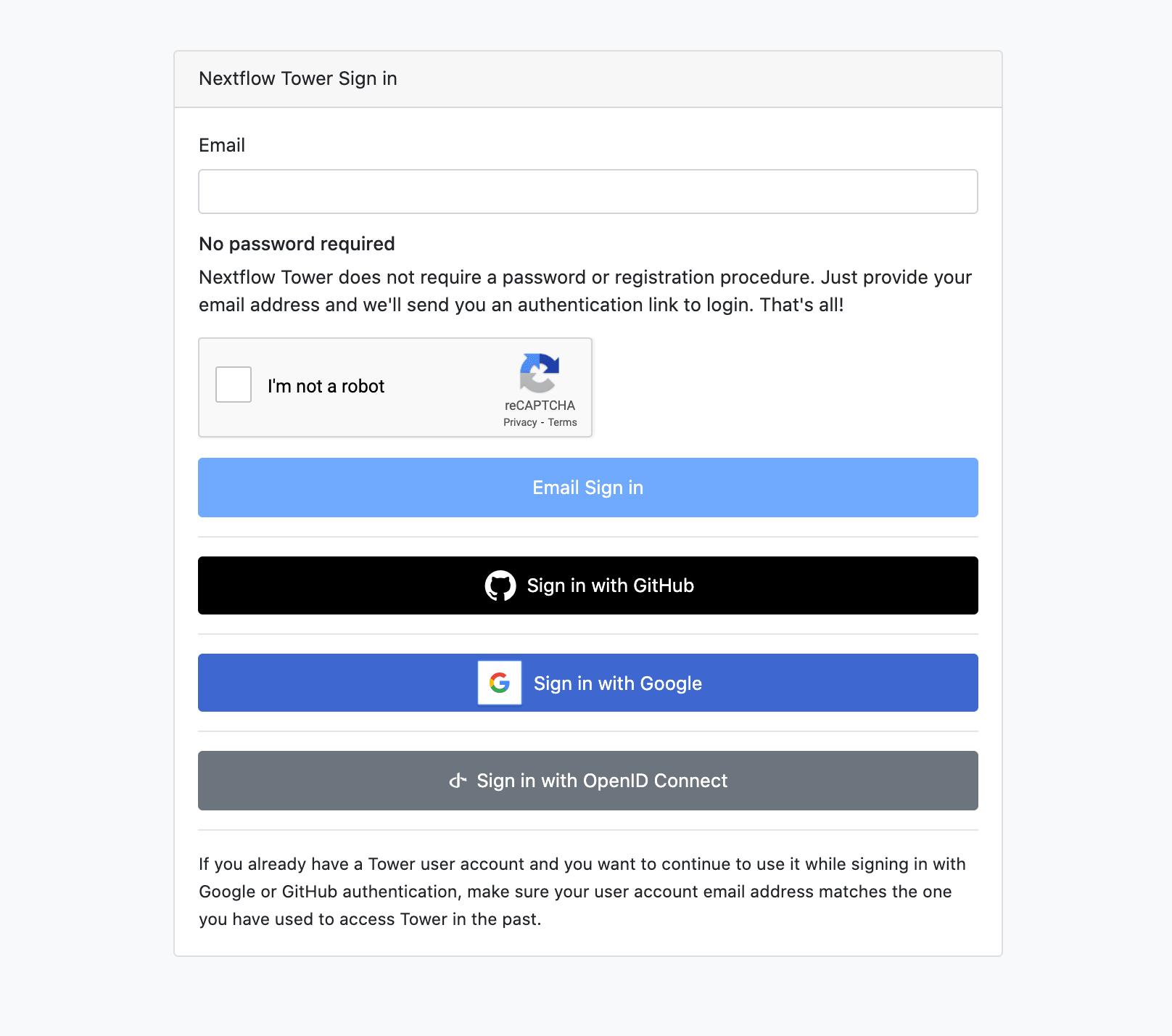
Finally, we have added a new login option to Nextflow Tower, allowing users to sign-in with their Google account. This will improve the experience and further broaden the audience outside of GitHub registered users.
Conclusion
Today's release addresses some of the most recurrent requests of Tower users, but there is still a lot to do. It is the input of scientists, developers and engineers that drives the Tower roadmap forward. From the team at Seqera, we'd like to thank you for every discussion, issue, DM and PR over the last 12 months. We're excited about the continued engagement and look forward to working together over the next 12 months as we shape the future of highly scalable and portable data analysis pipelines with Nextflow.
If your organization is interested in deploying Tower in your cloud or on-prem environment, please reach out to us at info@seqera.io, and we would be happy to discuss your requirements.
